Mammals
Inferring the mammal tree: species-level sets of phylogenies for questions in ecology, evolution, and conservation
Nathan S. Upham, Jacob A. Esselstyn, and Walter Jetz.
PLoS Biology 2019, https://doi.org/10.1371/journal.pbio.3000494
Explore the new mammal tree of life!
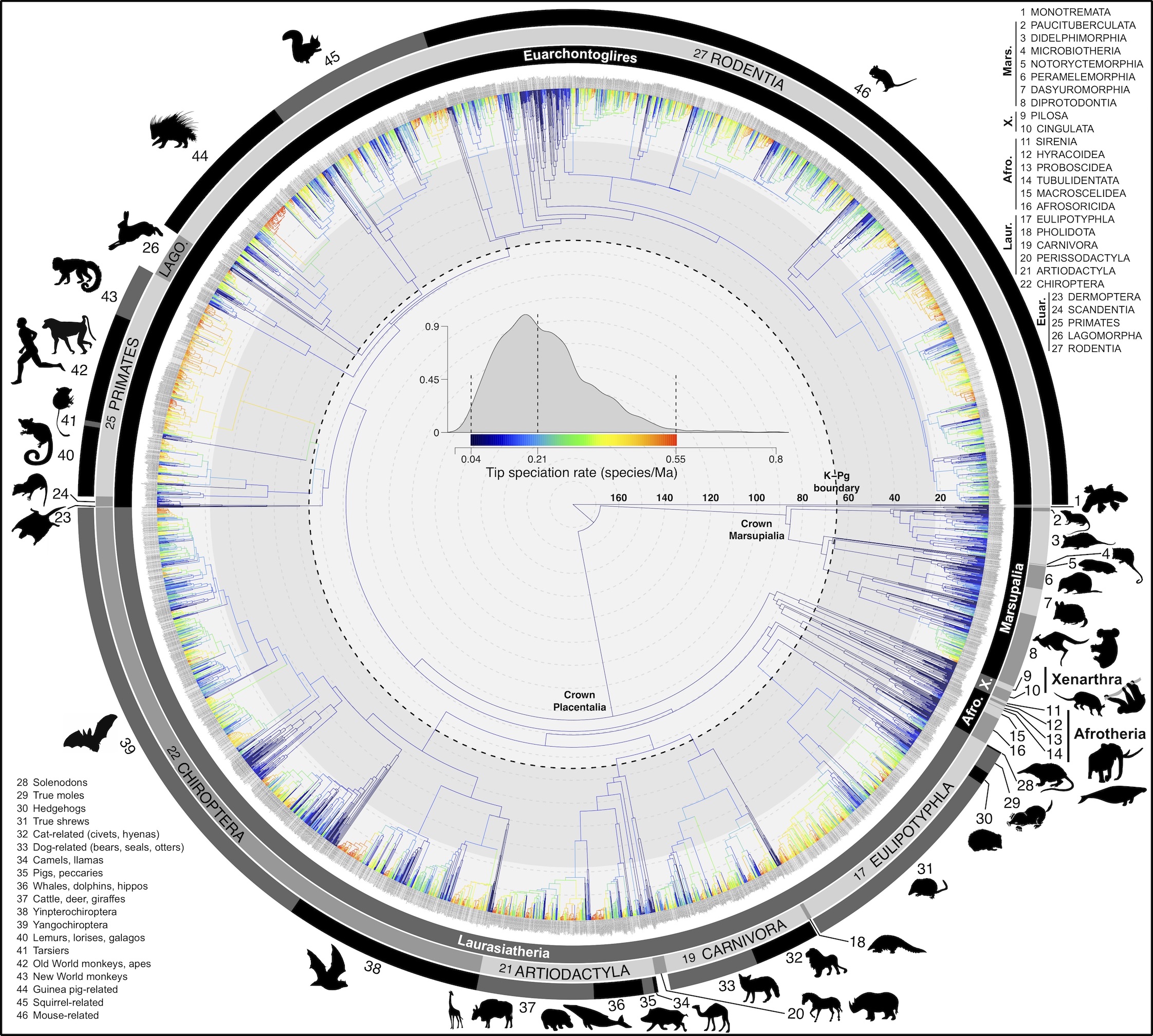
Hosted here on OneZoom is the consensus tree of 5,911 species of mammals, reconstructed using probabilistic inference of molecular and fossil data for the first time (see full comparison to previous methods here).
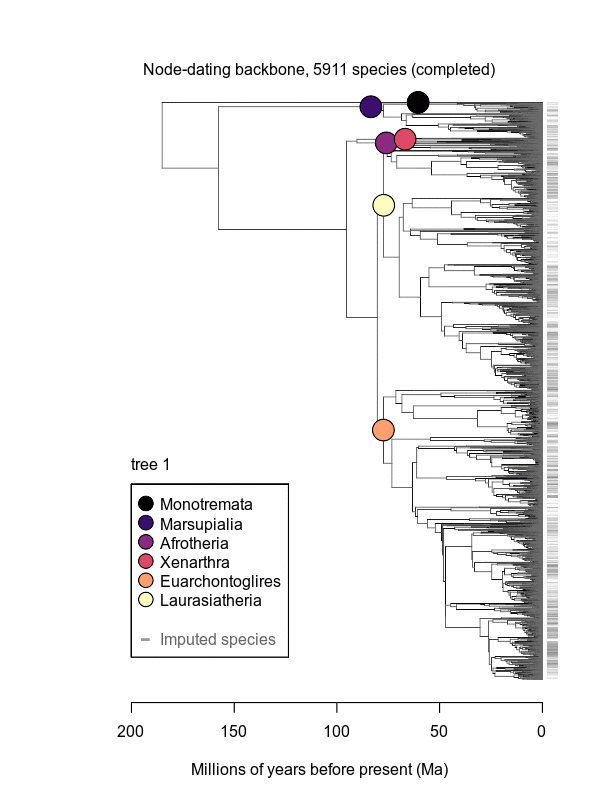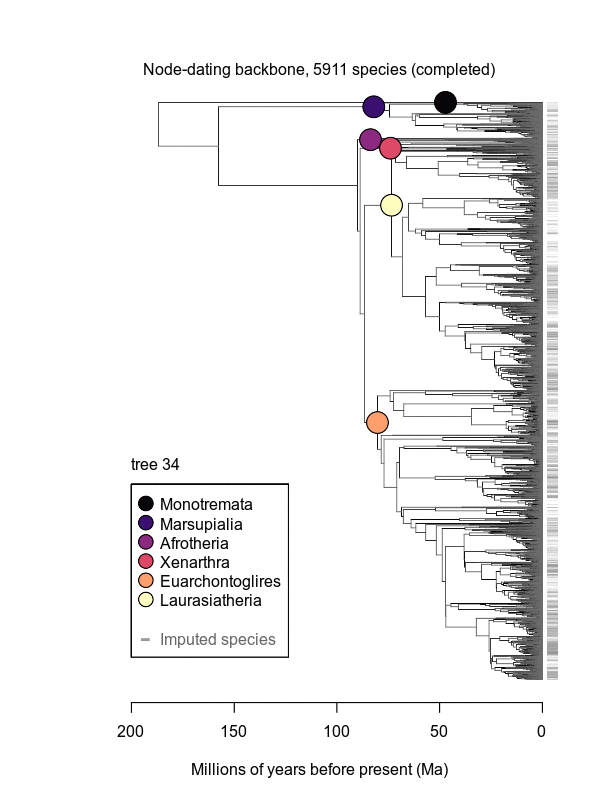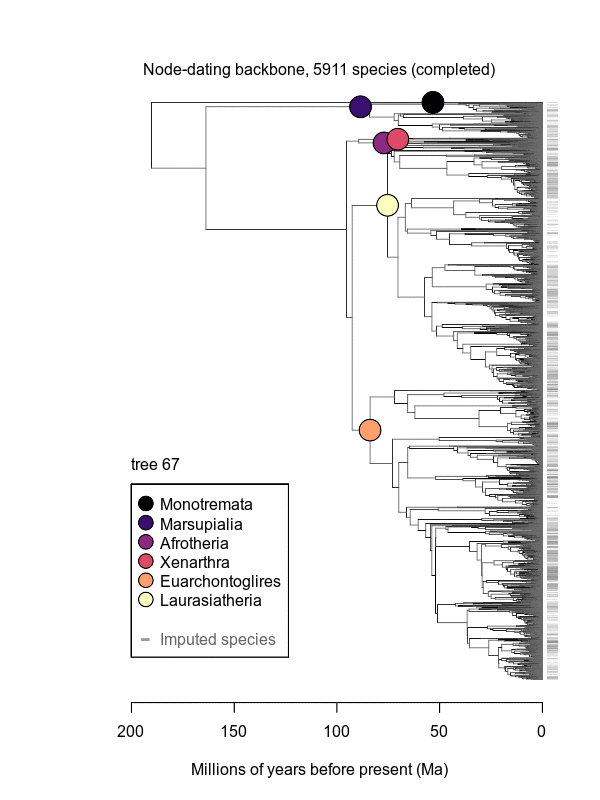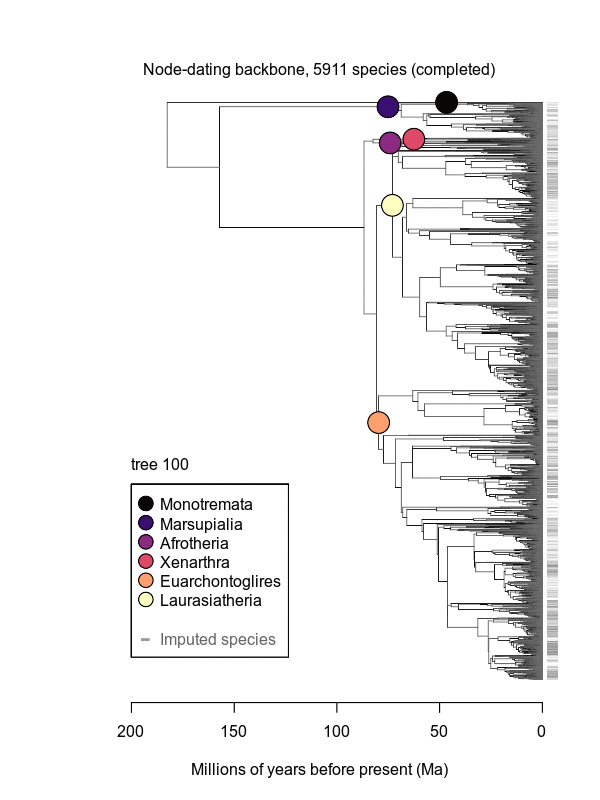
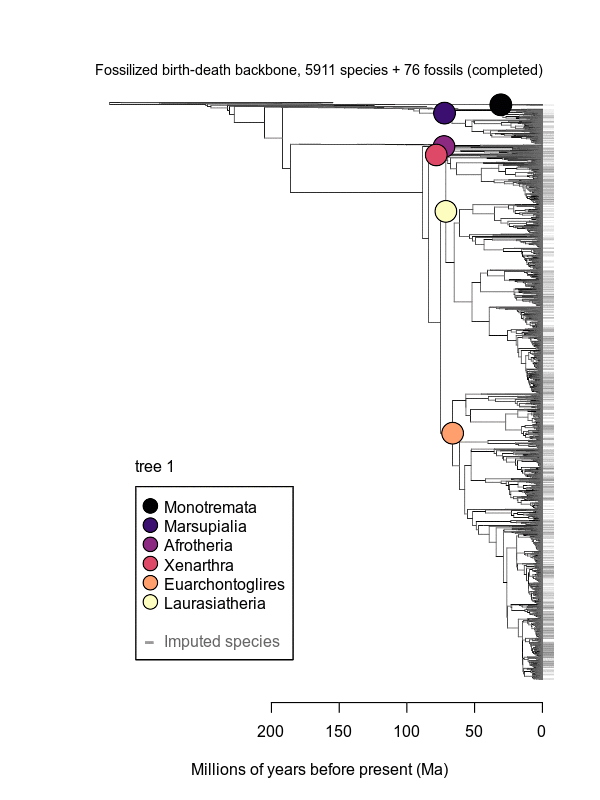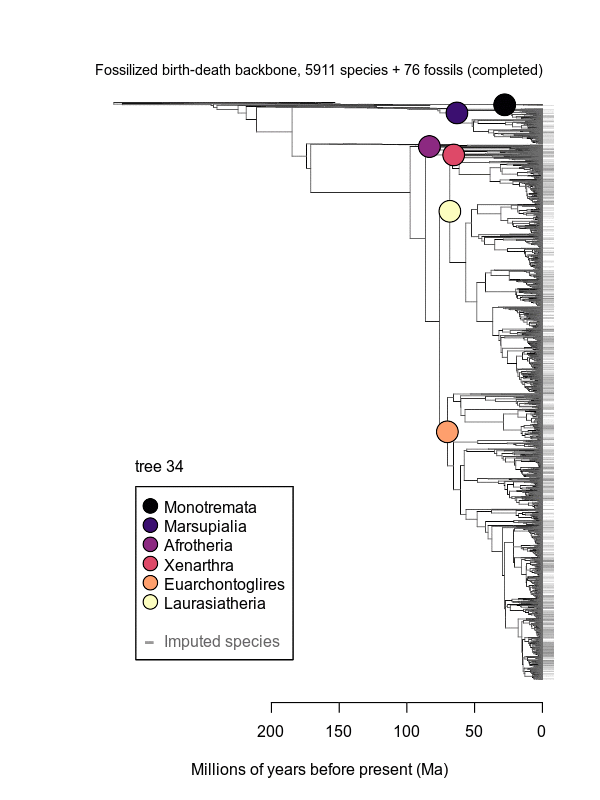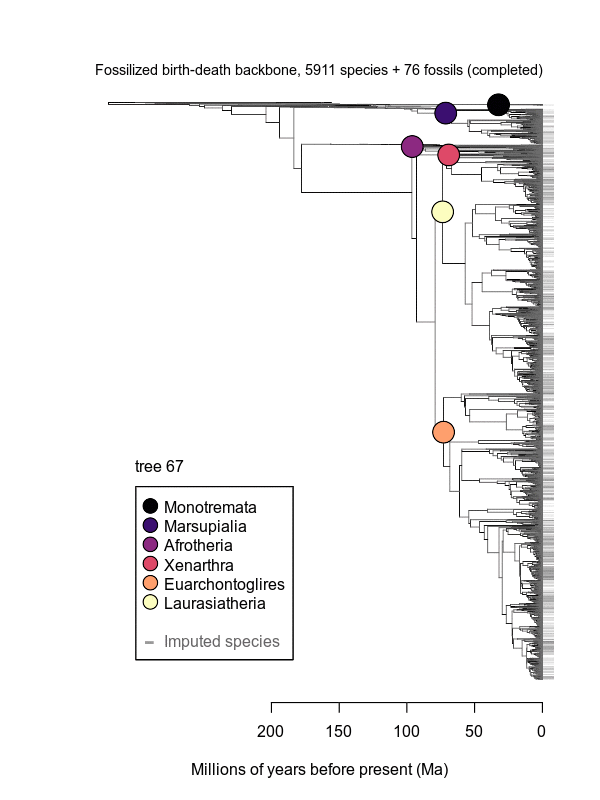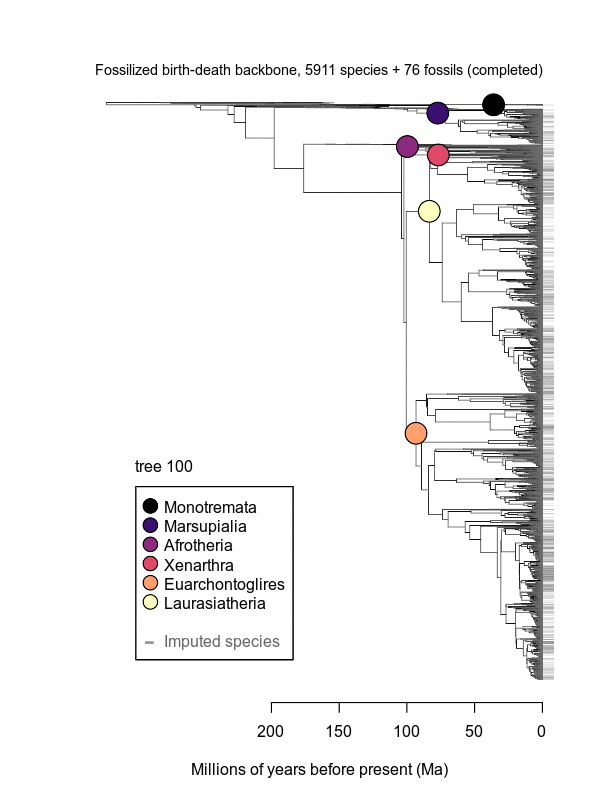
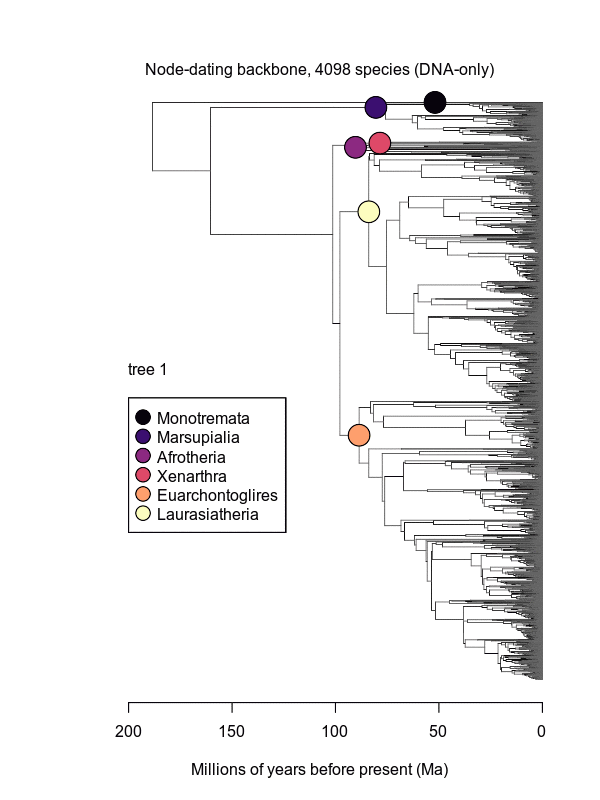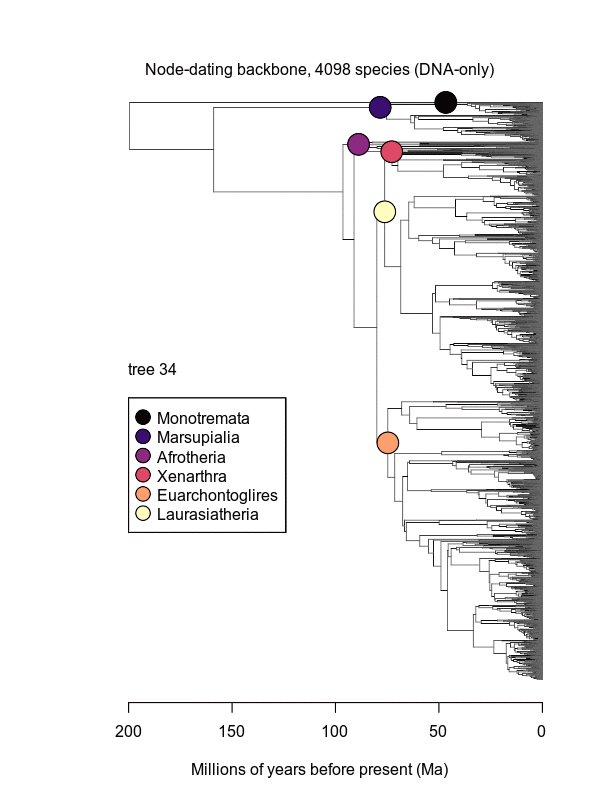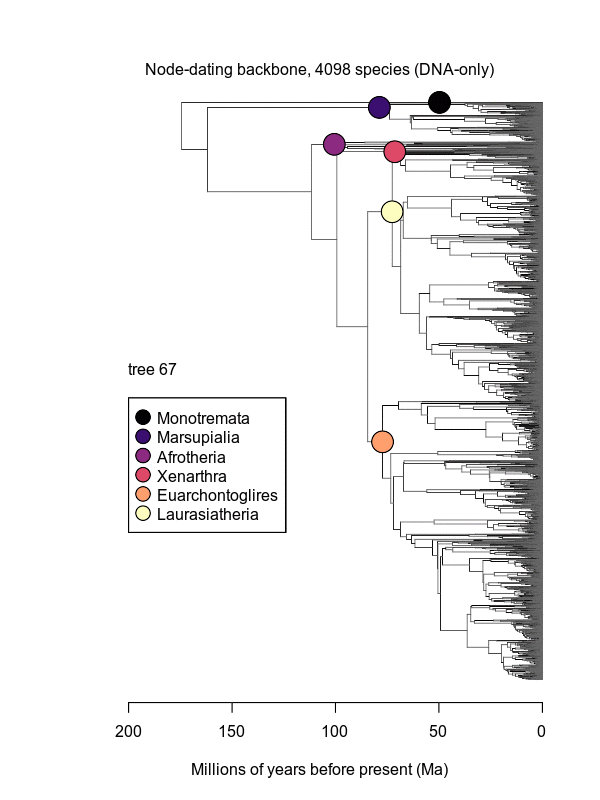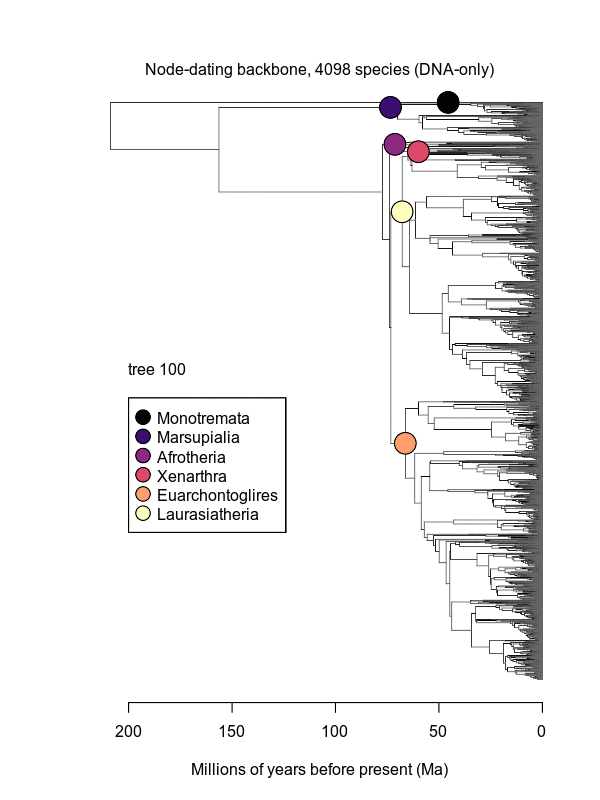
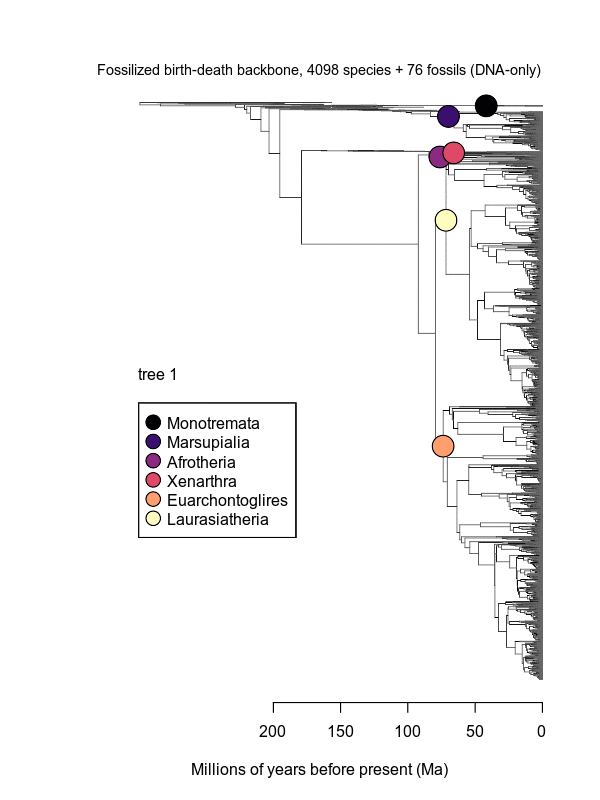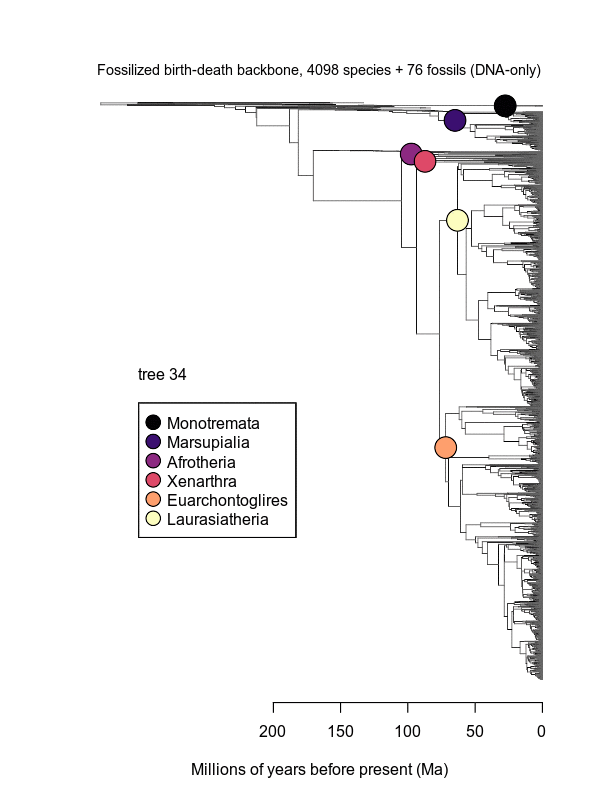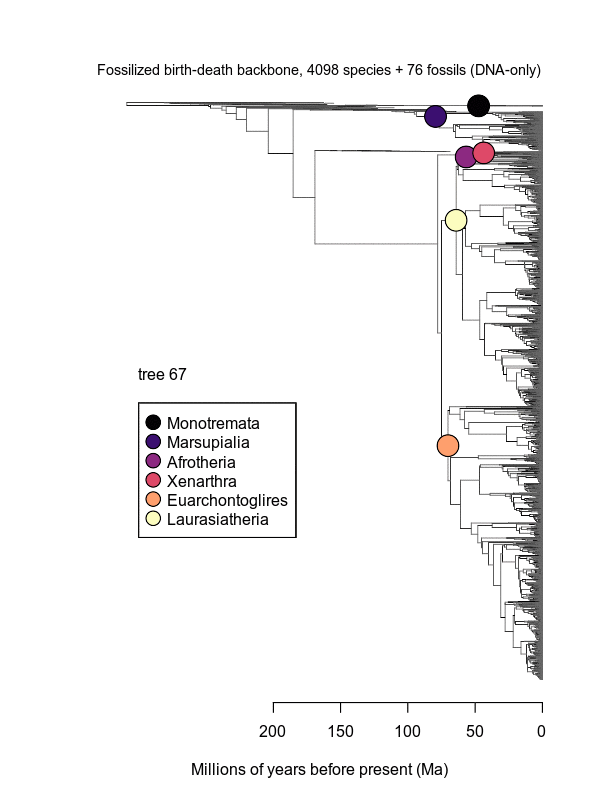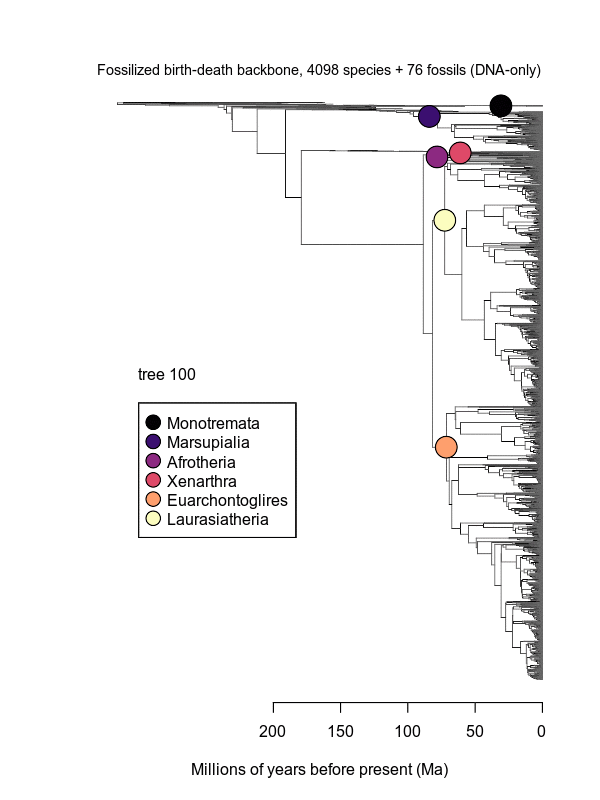
Movies of the credible sets of Mammalia trees (sample of 100 trees each of 10,000 total)
Shown is the evolutionary uncertainty propagated into each credible set of trees, consisting of both age and topological variation. The gray bars on the right show the location of the 1813 imputed species (i.e., those missing DNA sequences, of 5911 species total).
 |
 |
|---|---|
 |
 |
Citation: Upham, N. S., J. A. Esselstyn, and W. Jetz. 2019. Inferring the mammal tree: species-level sets of phylogenies for questions in ecology, evolution, and conservation. PLOS Biology. https://doi.org/10.1371/journal.pbio.3000494
Abstract: Big, time-scaled phylogenies are fundamental to connecting evolutionary processes to modern biodiversity patterns. Yet inferring reliable phylogenetic trees for thousands of species involves numerous trade-offs that have limited their utility to comparative biologists. To establish a robust evolutionary timescale for all ~6000 living species of mammals, we developed credible sets of trees that capture root-to-tip uncertainty in topology and divergence times. Our ‘backbone-and-patch’ approach to tree-building applies a newly assembled 31-gene supermatrix to two levels of Bayesian inference: (i) backbone relationships and ages among major lineages, using fossil node- or tip-dating; and (ii) species-level ‘patch’ phylogenies with non-overlapping in-groups that each correspond to one representative lineage in the backbone. Species unsampled for DNA are either excluded (‘DNA-only’ trees) or imputed within taxonomic constraints using branch lengths drawn from local birth-death models (‘completed’ trees). Joining time-scaled patches to backbones results in species-level trees of extant Mammalia with all branches estimated under the same modeling framework, thereby facilitating rate comparisons among lineages as disparate as marsupials and placentals. We compare our phylogenetic trees to previous estimates of mammal-wide phylogeny and divergence times, finding that (i) node ages are broadly concordant among studies, and (ii) recent (tip-level) rates of speciation are estimated more accurately in our study than in previous ‘supertree’ approaches where unresolved nodes led to branch length artifacts. Credible sets of mammalian phylogenetic history are now available for download at http://vertlife.org/phylosubsets, enabling investigations of long-standing questions in comparative biology.
Supplementary files:
-
Dryad of full data: https://doi.org/10.5061/dryad.tb03d03
-
Taxonomic subsetting and download of mammal trees: http://verlife.org/phylosubsets
-
All code for running analyses and plotting figures: https://github.com/n8upham/MamPhy_v1
-
MCC consensus trees of the DNA-only distributions:
– Node-dated exponential, 4098 species, backbone estimated with 17 node calibrations: pdf file (150 inches long); Nexus file
– Fossilized birth-death, 4098 species + 76 fossil tips, backbone topology as in Zhou et al. (2013): pdf file (150 inches long); Nexus file
-
MCC consensus trees of the completed trees (FOR DISPLAY ONLY): Note that completed trees are inappropriate for traditional single-tree analyses, since they contain DNA-missing species that randomly vary in topological position within taxonomic constraints (genus or family) across the credible set of trees. Thus, the completed trees must be analyzed as a sample of 100 or 1000 trees; any single tree is not a meaningful representation of the unknown phylogenetic placement of these imputed species. Fig 1 in our paper is an example of plotting the MCC completed tree for display purposes, but samples of trees were always analyzed to test hypotheses. See the “Recommended uses” section of our PLOS Biology article for more discussion of this topic.
– Node-dated exponential, 5911 species, backbone estimated with 17 node calibrations: Nexus file
– Fossilized birth-death, 5911 species + 76 fossil tips, backbone topology as in Zhou et al. (2013): Nexus file
Ecological causes of speciation and species richness in the mammal tree of life
Nathan S. Upham, Jacob A. Esselstyn, and Walter Jetz.
bioRxiv https://doi.org/10.1101/504803v3
-
On Twitter here
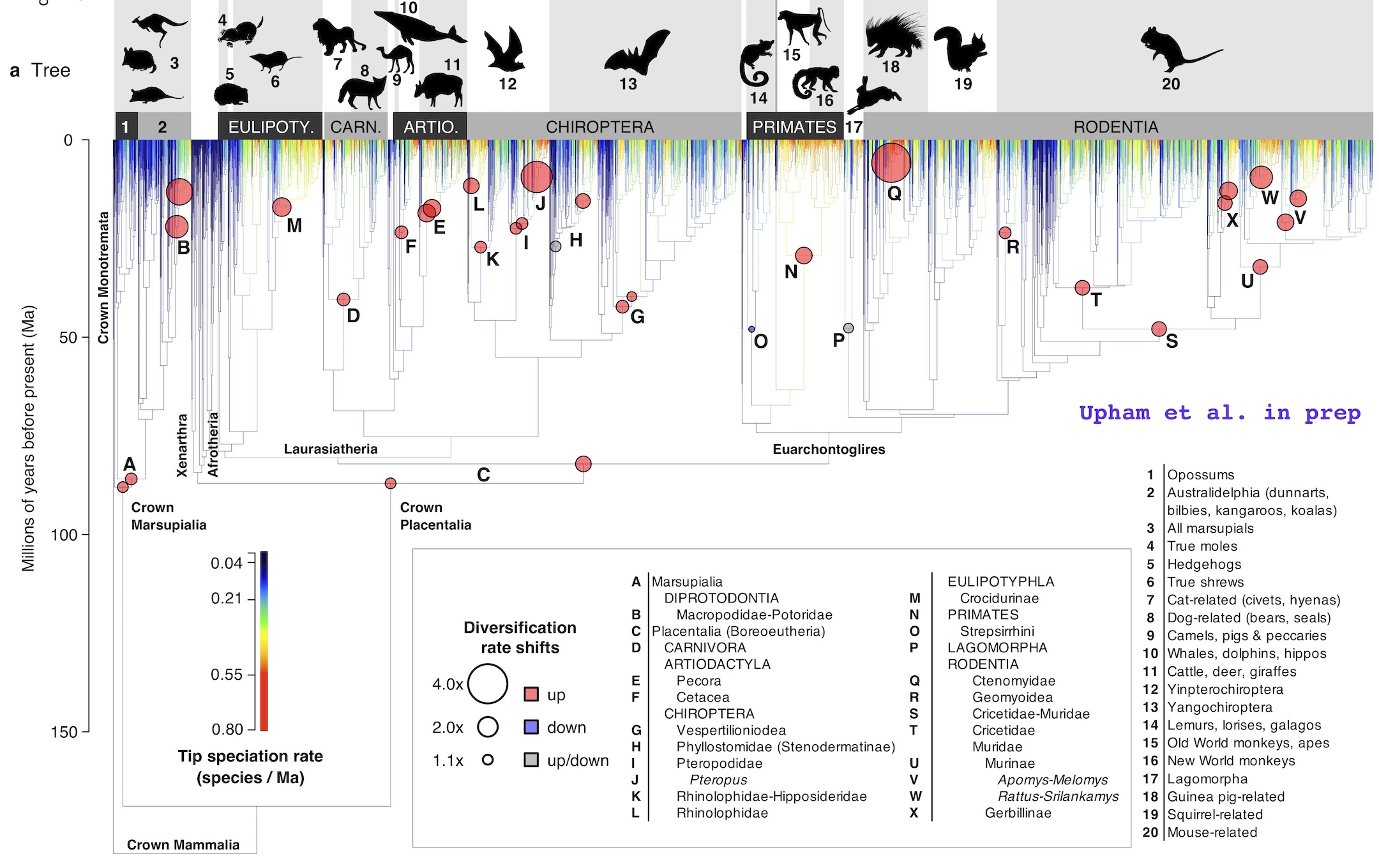
Abstract: Biodiversity is distributed unevenly from the poles to the equator, and among branches of the tree of life, yet how those patterns are related is unclear. We investigated global speciation-rate variation across crown Mammalia using a novel time-scaled phylogeny (N=5,911 species, ~70% with DNA), finding that trait- and latitude-associated speciation has caused uneven species richness among groups. We identify 24 branch-specific shifts in net diversification rates linked to ecological traits. Using time-slices to define clades, we show that speciation rates are a stronger predictor of clade richness than age. Mammals that are low dispersal or diurnal diversify the fastest, indicating roles for geographic and ecological speciation, respectively. Speciation is slower in tropical than extra-tropical lineages, consistent with evidence that longer tropical species durations underpin the latitudinal diversity gradient. These findings juxtapose modes of lineage diversification that are alternatively turnover-based, and thus non-adaptive, or persistence-based as associated with resource adaptations.
Taxonomy
The Mammal Diversity Database
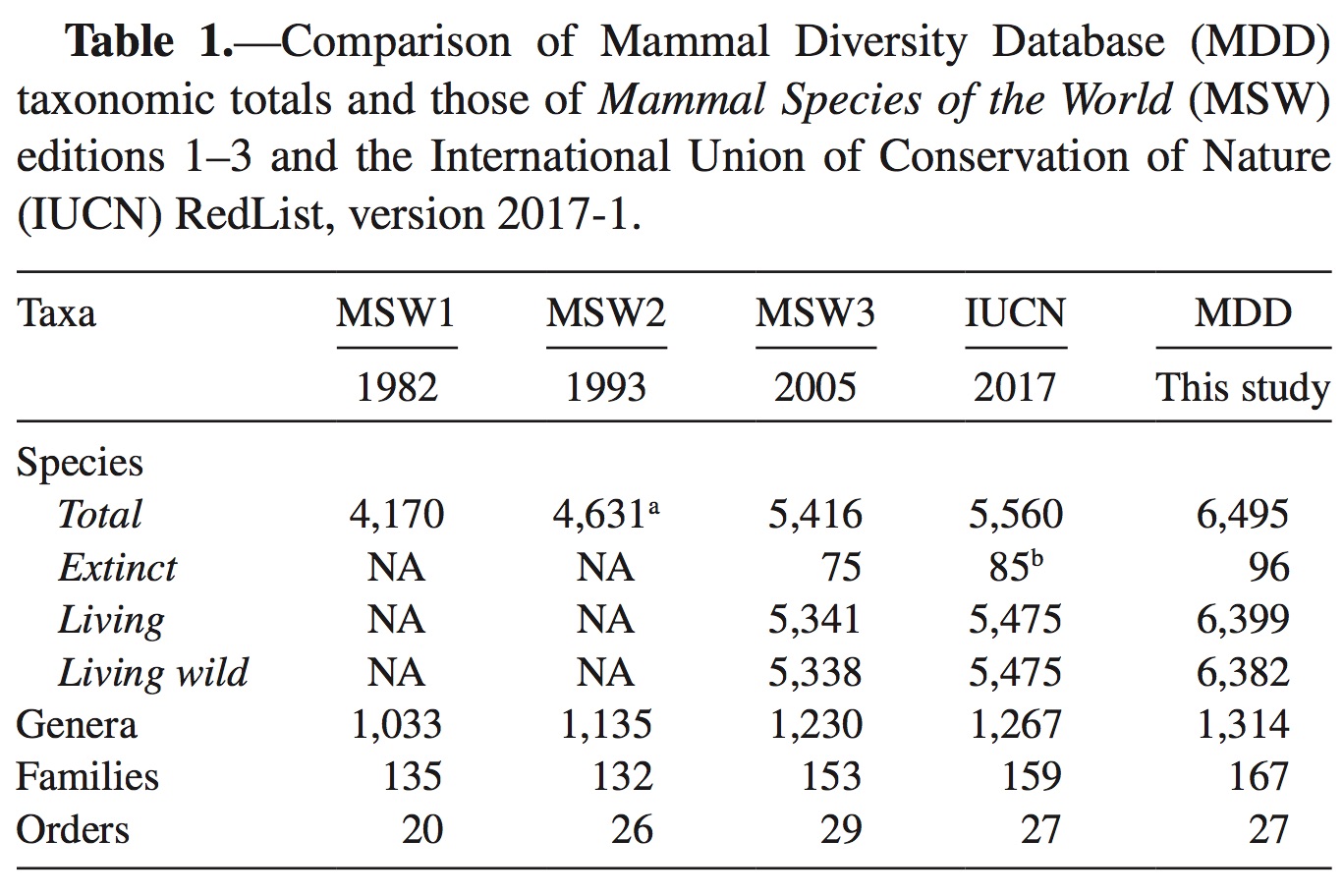
Citation: Burgin, C. J., J. P. Colella, P. L. Kahn, and N. S. Upham. 2018. How many species of mammals are there? Journal of Mammalogy 99:1–14. https://academic.oup.com/jmammal/article/99/1/1/4834091
Logistical and planning support for this work came from the NSF Vertlife Terrestrial grant with development commissioned by the American Society of Mammalogists.
The ASM Biodiversity Committee stewards the Mammal Diversity Database, an updatable and online database of mammal taxonomic and biodiversity information. This database aims to serve the global scientific community by providing the latest information on species-level and higher taxonomic changes, thereby promoting more rigorous study of mammalian biodiversity worldwide. The initial objective for this online database is to aggregate, curate, and compile new citations on species descriptions and taxonomic revisions into regular releases that are downloadable in comma-delimited format. Downstream goals include expanded hosting of ecological, trait, and taxonomic data, and an online forum for discussing mammalian taxonomy and systematics.
Traits
EltonTraits 1.0: Species-level foraging attributes of the world’s birds and mammals
Citation: Hamish Wilman, Jonathan Belmaker, Jennifer Simpson, Carolina de la Rosa, Marcelo M. Rivadeneira, and Walter Jetz. 2014. EltonTraits 1.0: Species-level foraging attributes of the world’s birds and mammals. Ecology 95:2027. http://dx.doi.org/10.1890/13-1917.1





.svg.png)